Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) uses multiple molecular mechanisms that allow for efficient infection into human cells and block activation of certain immunological response systems. Researcher from the Jagiellonian University – Dr
The methylation of the RNA Cap at 2′O-ribose is a specific type of camouflage that hides viral transcript in eukaryotic cells. This small modification prevents activation of certain pathways of hosts’ innate immune response linked to the production of interferons.
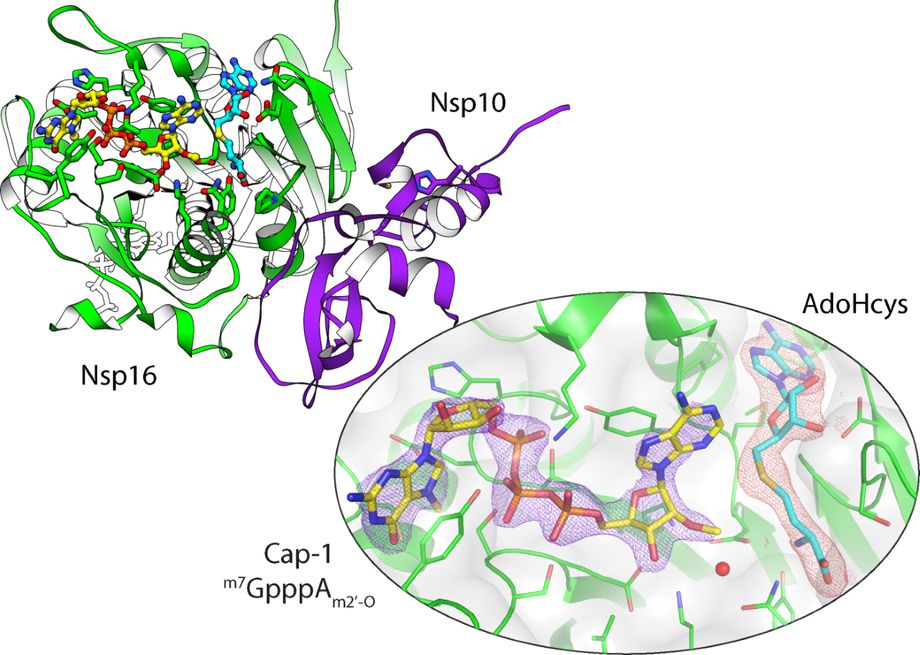
Transcripts are fragments of RNA that contain coding sequences – information that is used during translation to produce proteins. Modifications of these coding RNAs are factors which affect the degradation of transcripts, and modulate the efficiency of protein biosynthesis. After invading host cells, SARS-CoV-2 replicates the genetic material and translates viral proteins using the host protein translation machinery. The Nsp10/16 is a heterodimer of SARS-CoV-2 proteins that has 2′-O-methyltransferase activity. Using serial crystallography the crystal structure of Nsp10/16 was uncovered in a complex with methylated RNA Cap. The transfer of methyl group by Nsp10/16 is dependent on AdoMet, which serves as a methyl donor. The products of the reaction catalysed by Nsp10/16 are AdoHcys and 2′O methylated cap of the transcript – the structure called Cap-1. The occurrence of Cap-1 at the end of viral transcript allows SARS-CoV-2 to trick human innate response systems that recognize invading/exogenous RNA, and prevent the activation of interferon-dependent immunological response.
The article entitled “2′-O methylation of RNA cap in SARS-CoV-2 captured by serial crystallography” has been published in the prestigious journal Proceedings of the National Academy of Sciences of the United States of America. The article describes 4 crystal structures of Nsp10/16 complexes with substrates and products of methyl group transfer. This publication is the result of collaborative work of scientists from Argonne National Laboratory, the University of Chicago, the University of Northwestern, and the Jagiellonian University. Dr Wilamowski performed the experimental part of the research during his postdoctoral scholarship at the University of Chicago as part of the team lead by Prof. Andrzej Joachimiak. The crystallographic measurements were performed at the synchrotron facility – Advanced Photon Source (APS) in the United States. Ten of thousands of crystal diffraction images collected during a serial crystallography experiment were analysed using the Theta supercomputer at Argonne National Laboratory. The methodology of serial crystallography allows scientists to collect data from thousands of microcrystals at room-temperature, and therefore reduce the effect of radiation damage on determined protein structures.
Despite an advanced global vaccination programme against SARS-CoV-2 the effective treatment of COVID-19 patients requires the discovery of a drug. Small molecules that affect activity of the Nsp10/16 complex and inhibit 2′-O methylation of viral transcripts are good candidates for intelligent drug design using molecular modelling. The inhibitory molecule of Nsp10/16 methyltransferase could make SARS-CoV-2 sensitive to early activation of human innate immune response.